Last modified July 27, 2004.
Back to main page.
data;Then specify the number of variable residues:
param num_posn := p ;and the total number of rotamers:
param num_nodes := n ;Specify how many rotamers are in each position. The first column gives the position, the second gives the number of rotamers in that position. The sum of the position sizes must equal n.
param posn_size := 1 6 2 81 3 27 ... ;
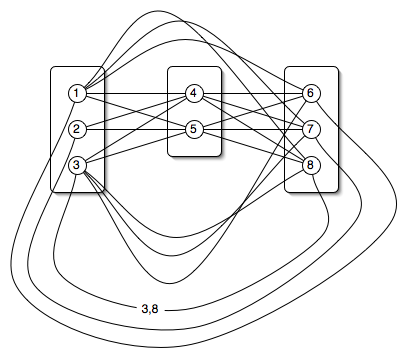 Specify the self-energies.
Each rotamer is given a number between 1 and
n. The rotamers of the first position are labeled 1 to the number of
rotamers for the first residues. The rotamers in the second position are
labeled with numbers starting right after the rotamers in the first position.
The first column is the rotamer number, the
second is the self-energy (in whatever units you want).
Specify the self-energies.
Each rotamer is given a number between 1 and
n. The rotamers of the first position are labeled 1 to the number of
rotamers for the first residues. The rotamers in the second position are
labeled with numbers starting right after the rotamers in the first position.
The first column is the rotamer number, the
second is the self-energy (in whatever units you want).
Specify the pairwise energies. The first two columns are rotamer numbers involved in the interaction. The third column is the interaction energy (in whatever units you want). The number in the first column must be less than the number in the second column. Don't specify pairs that have zero energy between them.param costV := 1 3.254 2 4.560 3 6.721 ... ;
Optionally, specify the number of solutions you want. This option only has an effect for integer programs. It is ignored for linear programs.param costE := 1 5 4.23 1 6 -3.2 3 5 -10 ... ;
param iterations := i ;Comments may be specified anywhere in the file by starting a line with a "#" character. The comment extends to the end of the line. Whitespace and, except in comments, newlines are ignored so you can format the file in a way that is convenient.
An example input file: 1igd.native.dat.gz.
The entire dataset in this format is available here.
ampl: model cks_lp.mod;Load the particular problem you want to solve:
ampl: data 1igd.native.dat;Then execute the Ampl script that matches the model you loaded:
ampl: commands cks_lp.ampl;To save typing, the above process is wrapped into two simple scripts so you can say:
% solve_cks_lp 1igd.native.dat > 1igd.native.lpsolwhere "%" is your system's prompt. You should edit these scripts and change the value of the LP variable to point to the directory containing the .mod and .ampl files. Also, Ampl should be added to your PATH.
% solve_cks_ilp 1igd.native.dat > 1igd.native.lpsol
The remaining sections of the file are delimited by lines beginning with "#" and a particular phrase. The order of the sections is fixed and the first one will be the energy of the optimal solution:492002 variables, all linear 50064 constraints, all linear; 1032460 nonzeros 1 linear objective; 242837 nonzeros. CPLEX 7.1.0: primal dualopt absmipgap=0 mipgap=1e-9 timing=1 Times (seconds): Input = 3.7 CPU 4.8 Wall Solve = 44 CPU 45 Wall Output = 0.36 CPU 0.43 Wall CPLEX 7.1.0: optimal solution; objective -765.4709674 22929 dual simplex iterations (13037 in phase I)
The second section gives the problem dimensions: the first line gives the total number of rotamers; the second line gives the number of variables residues.# energy -765.4709673910816
The next section gives the actual values for the node variables. If the solution is integral, it is these variables that specify which rotamer was chosen (1 = chosen, 0 = not chosen). The first column gives the rotamer number. The second indicates whether it was chosen. Lines for non-chosen rotamers may be omitted. For example:# problem size 1523 85
# primal vertex variables 1 1 2 0 4 0 ...
An example LP solution file: 1igd.native.lpsol.
(LP) option cplex_options 'primal dualopt timing=1';We've found that primal dualopt is the fastest solver option. timing=1 prints how long CPLEX spent on the problem. absmipgap=0 mipgap=1e-9 are the strictest settings for the ILP. They force CPLEX to return the optimal solution, rather than some close approximation to it.
(ILP) option cplex_options 'primal dualopt absmipgap=0 mipgap=1e-9 timing=1';
These options have been tested with CPLEX 7.1 and 7.5. Other versions may require slightly different options.
visitors since July 27, 2004.
Last modified July 27, 2004.
Back to main page.